PBMC model comparison#
Audience:
Researchers who want a quick baseline benchmark before building a custom model.
Prerequisites:
Install
scdlkit[tutorials].Know the Scanpy PBMC quickstart notebook first.
Learning goals:
Compare
PCA,autoencoder,vae, andtransformer_aeon the same PBMC workflow.Inspect the metrics table, runtime, and saved comparison plot.
Understand whether deep-learning baselines improve on a classical reference.
Out of scope:
full biological interpretation
claiming one benchmark chart settles model choice for every dataset
Install:
python -m pip install "scdlkit[tutorials]"
Published tutorial status
This page is a static notebook copy published for documentation review. It is meant to show the exact workflow and outputs from the last recorded run.
Last run date (UTC):
2026-03-27 09:24 UTCPublication mode:
static executed tutorialExecution profile:
publishedArtifact check in this sync:
passedSource notebook:
examples/compare_models_pbmc.ipynb
Outline#
Load PBMC data with Scanpy.
Detect the runtime device.
Choose the notebook profile.
Run the deep-learning baselines with aligned CPU-friendly settings.
Add
PCAas a classical reference baseline.Review scalar metrics and runtime.
Inspect saved plots and UMAP-style qualitative outputs.
from __future__ import annotations
from pathlib import Path
from time import perf_counter
import numpy as np
import pandas as pd
import scanpy as sc
import torch
from IPython.display import display
from scipy import sparse
from sklearn.decomposition import PCA
from scdlkit import compare_models
from scdlkit.evaluation.metrics import reconstruction_metrics, representation_metrics
from scdlkit.visualization.compare import plot_model_comparison
DATA_PATH = Path("examples/data/pbmc3k_processed.h5ad")
OUTPUT_DIR = Path("artifacts/pbmc_compare")
OUTPUT_DIR.mkdir(parents=True, exist_ok=True)
device_name = "cuda" if torch.cuda.is_available() else "cpu"
print(f"Using device: {device_name}")
def save_umap_from_latent(
adata,
latent,
output_path,
*,
label_key="louvain",
use_rep="X_quality_latent",
):
plot_adata = adata.copy()
plot_adata.obsm[use_rep] = latent
sc.pp.neighbors(plot_adata, use_rep=use_rep)
sc.tl.umap(plot_adata, random_state=42)
fig = sc.pl.umap(plot_adata, color=label_key, return_fig=True, frameon=False)
fig.savefig(output_path, dpi=150, bbox_inches="tight")
return fig
Using device: cpu
TUTORIAL_PROFILE = "quickstart" # change to "full" for a longer run
PROFILE = {
"quickstart": {"epochs": 10, "batch_size": 128},
"full": {"epochs": 25, "batch_size": 128},
}[TUTORIAL_PROFILE]
TRANSFORMER_MODEL_KWARGS = {
"patch_size": 48,
"d_model": 64,
"n_heads": 2,
"n_layers": 1,
"decoder_hidden_dims": (128,),
}
print(f"Tutorial profile: {TUTORIAL_PROFILE}")
print(PROFILE)
print("Compact transformer settings:", TRANSFORMER_MODEL_KWARGS)
Tutorial profile: quickstart
{'epochs': 10, 'batch_size': 128}
Compact transformer settings: {'patch_size': 48, 'd_model': 64, 'n_heads': 2, 'n_layers': 1, 'decoder_hidden_dims': (128,)}
Load PBMC data#
The comparison uses the same PBMC dataset as the quickstart tutorial so the main variable is the baseline model, not the preprocessing story.
adata = sc.read_h5ad(DATA_PATH) if DATA_PATH.exists() else sc.datasets.pbmc3k_processed()
print(adata)
print("Label field used for comparison:", "louvain")
AnnData object with n_obs × n_vars = 2638 × 1838
obs: 'n_genes', 'percent_mito', 'n_counts', 'louvain'
var: 'n_cells'
uns: 'draw_graph', 'louvain', 'louvain_colors', 'neighbors', 'pca', 'rank_genes_groups'
obsm: 'X_pca', 'X_tsne', 'X_umap', 'X_draw_graph_fr'
varm: 'PCs'
obsp: 'distances', 'connectivities'
Label field used for comparison: louvain
Compare deep-learning baselines#
The benchmark keeps the task, label field, and training settings aligned so the model family is the main variable. The VAE uses a lighter KL term and the Transformer AE uses a compact CPU-friendly configuration so the comparison stays practical in docs and CI. We then add PCA as the classical reference baseline around those deep-learning rows.
from scdlkit import TaskRunner
base_shared_kwargs = {
"epochs": PROFILE["epochs"],
"batch_size": PROFILE["batch_size"],
"label_key": "louvain",
"device": "auto",
}
ae_result = compare_models(
adata,
models=["autoencoder"],
task="representation",
shared_kwargs=base_shared_kwargs,
output_dir=str(OUTPUT_DIR / "autoencoder"),
)
transformer_started_at = perf_counter()
transformer_runner = TaskRunner(
model="transformer_ae",
task="representation",
output_dir=str(OUTPUT_DIR / "transformer_ae"),
model_kwargs=TRANSFORMER_MODEL_KWARGS,
**base_shared_kwargs,
)
transformer_runner.fit(adata)
transformer_metrics = transformer_runner.evaluate()
transformer_runtime_sec = perf_counter() - transformer_started_at
transformer_row = pd.DataFrame([
{
"model": "transformer_ae",
"runtime_sec": transformer_runtime_sec,
**{
key: value
for key, value in transformer_metrics.items()
if isinstance(value, (int, float))
},
}
])
vae_started_at = perf_counter()
vae_runner = TaskRunner(
model="vae",
task="representation",
output_dir=str(OUTPUT_DIR / "vae"),
model_kwargs={"kl_weight": 1e-3},
**base_shared_kwargs,
)
vae_runner.fit(adata)
vae_metrics = vae_runner.evaluate()
vae_runtime_sec = perf_counter() - vae_started_at
vae_row = pd.DataFrame([
{
"model": "vae",
"runtime_sec": vae_runtime_sec,
**{
key: value
for key, value in vae_metrics.items()
if isinstance(value, (int, float))
},
}
])
deep_metrics = (
pd.concat([ae_result.metrics_frame, transformer_row, vae_row], ignore_index=True)
.sort_values("model")
.reset_index(drop=True)
)
deep_runners = {
**ae_result.runners,
"transformer_ae": transformer_runner,
"vae": vae_runner,
}
deep_metrics
| model | runtime_sec | mse | mae | pearson | spearman | silhouette | knn_label_consistency | ari | nmi | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | autoencoder | 4.532655 | 0.819260 | 0.402501 | 0.235012 | 0.128319 | 0.170472 | 0.868687 | 0.635203 | 0.758904 |
| 1 | transformer_ae | 7.465522 | 0.840910 | 0.407200 | 0.173332 | 0.074018 | 0.048607 | 0.689394 | 0.275133 | 0.397407 |
| 2 | vae | 3.496453 | 0.820916 | 0.399915 | 0.230418 | 0.115319 | 0.175449 | 0.898990 | 0.588731 | 0.770850 |
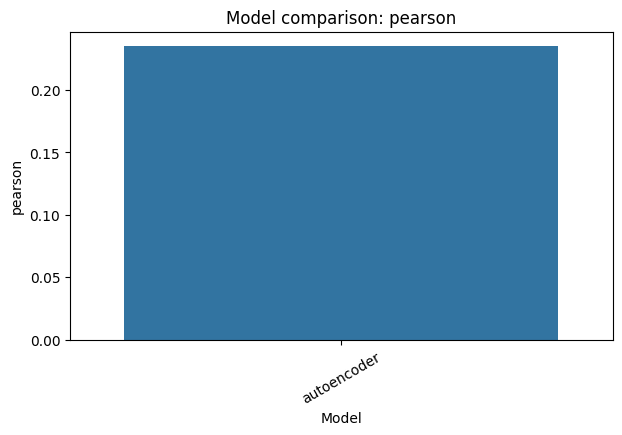
Add PCA as the classical reference baseline#
A baseline-first toolkit should not compare deep-learning models only against each other. Here we add a PCA row with the same representation and reconstruction metrics, then overwrite the saved comparison CSV and plot with the combined view.
x_matrix = adata.X.toarray() if sparse.issparse(adata.X) else np.asarray(adata.X, dtype="float32")
labels = pd.Categorical(adata.obs["louvain"].astype(str)).codes.astype(int)
pca_started_at = perf_counter()
pca = PCA(n_components=min(32, x_matrix.shape[0] - 1, x_matrix.shape[1]), random_state=42)
pca_latent = pca.fit_transform(x_matrix)
pca_reconstruction = pca.inverse_transform(pca_latent)
pca_runtime_sec = perf_counter() - pca_started_at
pca_metrics = reconstruction_metrics(x_matrix, pca_reconstruction)
pca_metrics.update(representation_metrics(pca_latent, labels, None))
pca_row = pd.DataFrame([
{
"model": "pca",
"runtime_sec": pca_runtime_sec,
**pca_metrics,
}
])
combined_metrics = (
pd.concat([pca_row, deep_metrics], ignore_index=True)
.sort_values("model")
.reset_index(drop=True)
)
combined_metrics.to_csv(OUTPUT_DIR / "benchmark_metrics.csv", index=False)
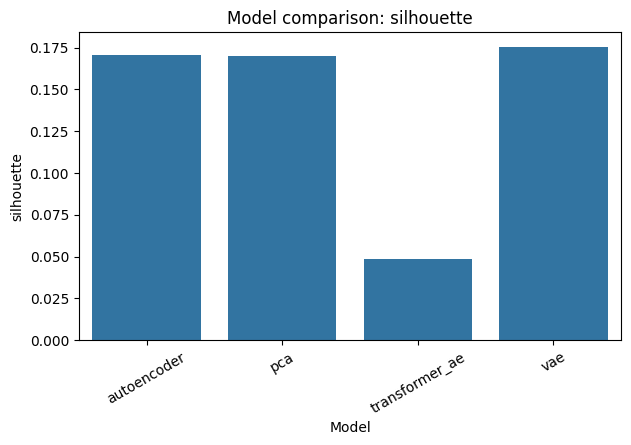
comparison_fig, _ = plot_model_comparison(combined_metrics, metric="silhouette")
comparison_fig.savefig(OUTPUT_DIR / "benchmark_comparison.png", dpi=150, bbox_inches="tight")
display(comparison_fig)
combined_metrics
| model | runtime_sec | mse | mae | pearson | spearman | silhouette | knn_label_consistency | ari | nmi | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | autoencoder | 4.532655 | 0.819260 | 0.402501 | 0.235012 | 0.128319 | 0.170472 | 0.868687 | 0.635203 | 0.758904 |
| 1 | pca | 0.126385 | 0.780000 | 0.416987 | 0.317878 | 0.169146 | 0.170164 | 0.948067 | 0.650855 | 0.798312 |
| 2 | transformer_ae | 7.465522 | 0.840910 | 0.407200 | 0.173332 | 0.074018 | 0.048607 | 0.689394 | 0.275133 | 0.397407 |
| 3 | vae | 3.496453 | 0.820916 | 0.399915 | 0.230418 | 0.115319 | 0.175449 | 0.898990 | 0.588731 | 0.770850 |

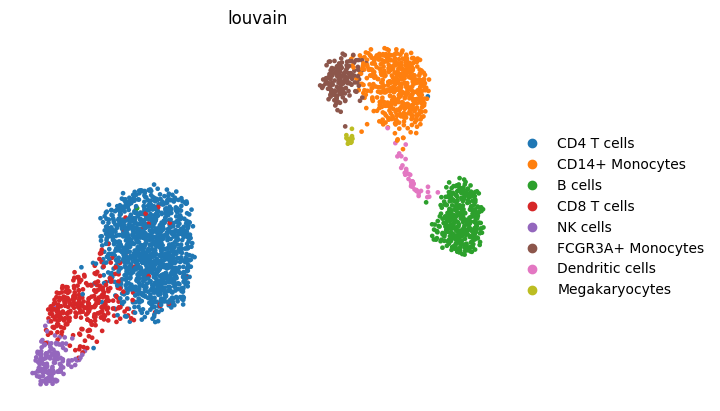
Inspect qualitative outputs#
Quantitative metrics should be matched with a qualitative check. Here the notebook saves one UMAP from the PCA reference and one UMAP from the strongest deep-learning baseline by silhouette.
pca_fig = save_umap_from_latent(
adata,
pca_latent,
OUTPUT_DIR / "pca_reference_umap.png",
use_rep="X_pca_reference",
)
display(pca_fig)
best_deep_row = deep_metrics.sort_values("silhouette", ascending=False).iloc[0]
best_deep_model = best_deep_row["model"]
best_runner = deep_runners[best_deep_model]
best_fig = save_umap_from_latent(
adata,
best_runner.encode(adata),
OUTPUT_DIR / "best_baseline_umap.png",
use_rep="X_best_baseline",
)
display(best_fig)
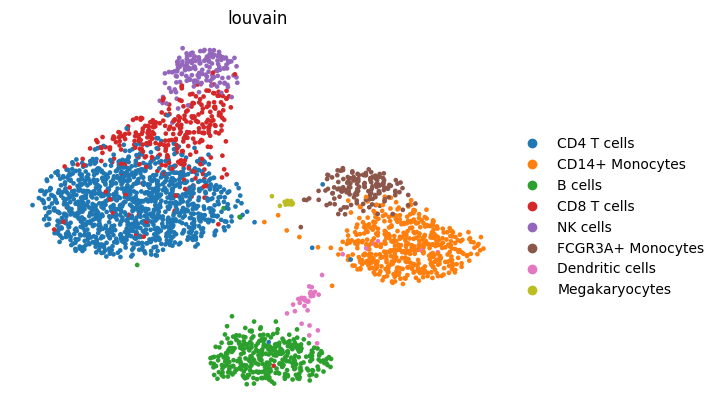


Generated outputs and interpretation#
The comparison writes its combined outputs to artifacts/pbmc_compare/.
Interpret the table in this order:
Check whether the deep-learning baselines beat
PCAon the representation metrics.Check how much runtime that gain costs.
Use the UMAP-style outputs to make sure the scalar metrics match the qualitative structure.
Do not treat a small win over
PCAas proof that a deeper model is always the right choice.
When PCA is enough:
the separation is already clean
the metric gain from deeper models is small
runtime matters more than a marginal score change
Recommended next tutorials:
downstream Scanpy after scDLKit when you want cluster interpretation
custom model extension when you want to beat the built-in baselines with your own model
best_overall = combined_metrics.sort_values("silhouette", ascending=False).iloc[0]
fastest_overall = combined_metrics.sort_values("runtime_sec", ascending=True).iloc[0]
print(f"Best model by silhouette: {best_overall['model']}")
print(f"Fastest model by runtime_sec: {fastest_overall['model']}")
Best model by silhouette: vae
Fastest model by runtime_sec: pca
combined_output_paths = {
"metrics_csv": str(OUTPUT_DIR / "benchmark_metrics.csv"),
"comparison_png": str(OUTPUT_DIR / "benchmark_comparison.png"),
"pca_umap_png": str(OUTPUT_DIR / "pca_reference_umap.png"),
"best_model_umap_png": str(OUTPUT_DIR / "best_baseline_umap.png"),
}
combined_output_paths
{'metrics_csv': 'artifacts/pbmc_compare/benchmark_metrics.csv',
'comparison_png': 'artifacts/pbmc_compare/benchmark_comparison.png',
'pca_umap_png': 'artifacts/pbmc_compare/pca_reference_umap.png',
'best_model_umap_png': 'artifacts/pbmc_compare/best_baseline_umap.png'}
