scDLKit#
scDLKit is moving from a baseline toolkit identity toward a publication-first research program with a software artifact attached to it.
Available now#
Today the public repo supports two main entrypoints:
stable baseline workflows through
TaskRunnerexperimental labeled annotation adaptation through
adapt_annotation(...)a static published annotation tutorial and status page for docs review
The current implemented scope is still narrower than the paper target:
stable deep-learning baselines for single-cell workflows
Scanpy handoff through
adata.obsmexperimental scGPT annotation adaptation on labeled human scRNA-seq
beyond-PBMC annotation evidence on human pancreas
Paper target#
The paper target is:
scDLKit is a minimal-code, AnnData-native framework for parameter-efficient adaptation and reproducible benchmarking of single-cell and spatial foundation models.
That target expands the repo in two directions:
model breadth:
scGPTscFoundationCellFMNicheformer
task breadth:
annotation
integration
perturbation
spatial
Use the roadmap when you want the full distinction between paper target and current implementation truth.
Main research task map#
Status: Pilot
Main question: Can scDLKit already support a credible low-code adaptation story on labeled human data?
Current implementation note:
The pilot currently runs on the experimental scGPT path only. The published
quickstart tutorial compares frozen_probe and head, while the heavier
annotation benchmark matrix extends to full fine-tuning, lora, adapter,
prefix_tuning, and ia3.
Status: Planned
Main question: Can adapted representations transfer across studies and batches under a standardized benchmark?
Status: Planned
Main question: Can the framework benchmark adaptation strategies on perturbation-response tasks?
Status: Planned
Main question: Can scDLKit support a real spatial pillar anchored by Nicheformer rather than a future placeholder?
Current entrypoints#
Stable baseline path#
import scanpy as sc
from scdlkit import TaskRunner
adata = sc.datasets.pbmc3k_processed()
runner = TaskRunner(
model="vae",
task="representation",
label_key="louvain",
device="auto",
epochs=20,
batch_size=128,
model_kwargs={"kl_weight": 1e-3},
)
runner.fit(adata)
adata.obsm["X_scdlkit_vae"] = runner.encode(adata)
Use this when you want the stable baseline workflow.
Related docs:
Experimental annotation path#
from scdlkit import adapt_annotation
runner = adapt_annotation(
adata,
label_key="cell_type",
output_dir="artifacts/scgpt_annotation",
)
runner.annotate_adata(adata)
runner.save("artifacts/scgpt_annotation/best_model")
Use this when you want the current low-code research-facing adaptation path.
Related docs:
Supporting workflows#
Workflow snapshots#
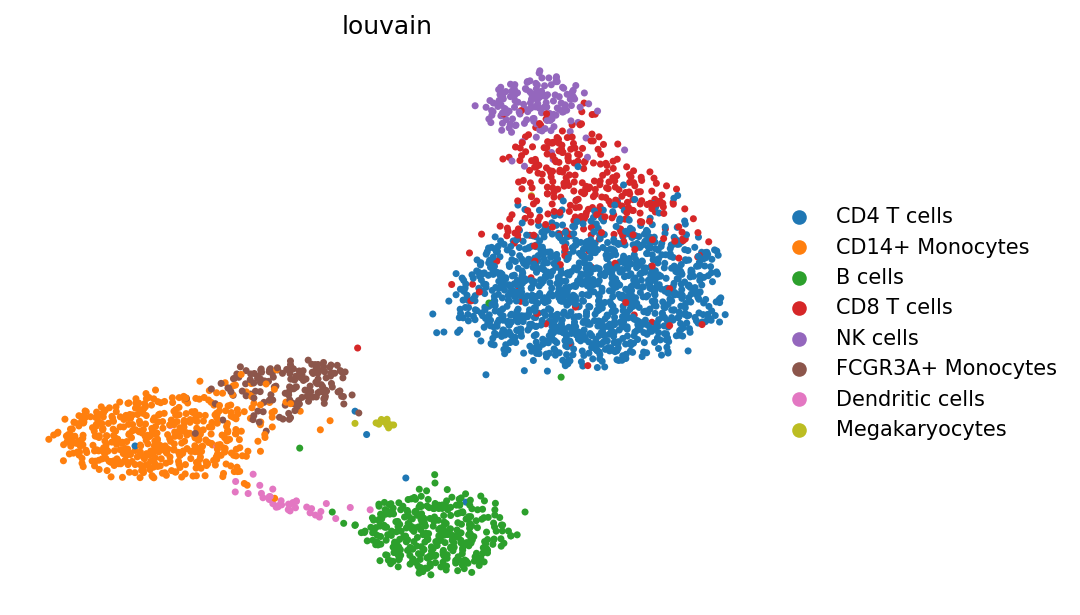
Quickstart embedding colored by the PBMC reference labels.#
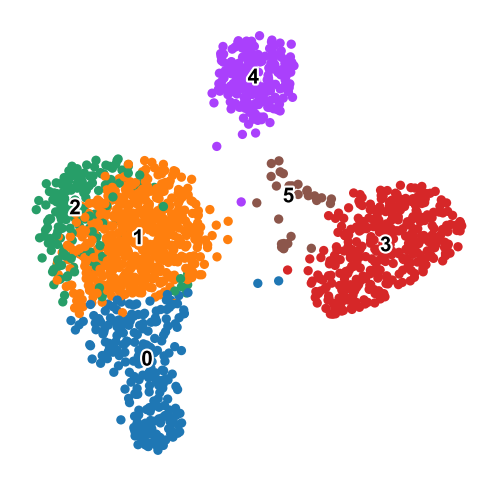
Leiden clustering on the same embedding after handing control back to Scanpy.#
Current scope#
Scanpy still owns raw-data preprocessing, QC, and most exploratory analysis.
scDLKit currently owns model training, evaluation, comparison, and output handoff.
the current public implementation is still gene-expression-first
the paper target is broader than the current implementation and must remain labeled as such
