PBMC classification baseline#
Audience:
Researchers who want a simple supervised baseline on a familiar single-cell dataset.
Prerequisites:
Install
scdlkit[tutorials].PBMC data available through Scanpy or the repository cache.
Learning goals:
Train
mlp_classifieron PBMC labels.Inspect accuracy and macro F1.
Plot and save a confusion matrix.
Keep the same notebook path on CPU or GPU with
device="auto".
Out of scope:
production-grade annotation
uncertainty calibration
large-scale cell-type labeling workflows
Install:
python -m pip install "scdlkit[tutorials]"
Published tutorial status
This page is a static notebook copy published for documentation review. It is meant to show the exact workflow and outputs from the last recorded run.
Last run date (UTC):
2026-03-27 09:24 UTCPublication mode:
static executed tutorialExecution profile:
publishedArtifact check in this sync:
passedSource notebook:
examples/classification_demo.ipynb
Outline#
Load PBMC data with Scanpy.
Detect the runtime device.
Choose the notebook profile.
Fit the classification baseline with
device="auto".Evaluate and save a report.
Plot a confusion matrix.
from __future__ import annotations
from pathlib import Path
import scanpy as sc
import torch
from IPython.display import display
from scdlkit import TaskRunner
DATA_PATH = Path("examples/data/pbmc3k_processed.h5ad")
OUTPUT_DIR = Path("artifacts/pbmc_classification")
OUTPUT_DIR.mkdir(parents=True, exist_ok=True)
device_name = "cuda" if torch.cuda.is_available() else "cpu"
print(f"Using device: {device_name}")
Using device: cpu
TUTORIAL_PROFILE = "quickstart" # change to "full" for a longer run
PROFILE = {
"quickstart": {"epochs": 15, "batch_size": 128},
"full": {"epochs": 30, "batch_size": 128},
}[TUTORIAL_PROFILE]
print(f"Tutorial profile: {TUTORIAL_PROFILE}")
print(PROFILE)
Tutorial profile: quickstart
{'epochs': 15, 'batch_size': 128}
adata = sc.read_h5ad(DATA_PATH) if DATA_PATH.exists() else sc.datasets.pbmc3k_processed()
print(adata)
print("Classification target:", "louvain")
AnnData object with n_obs × n_vars = 2638 × 1838
obs: 'n_genes', 'percent_mito', 'n_counts', 'louvain'
var: 'n_cells'
uns: 'draw_graph', 'louvain', 'louvain_colors', 'neighbors', 'pca', 'rank_genes_groups'
obsm: 'X_pca', 'X_tsne', 'X_umap', 'X_draw_graph_fr'
varm: 'PCs'
obsp: 'distances', 'connectivities'
Classification target: louvain
Train the classifier#
This is the same code path on CPU and GPU. The runner will select CUDA automatically when it is available.
Treat this notebook as a baseline classifier, not a production-ready cell-annotation system. Its job is to tell you whether a simple supervised MLP is already competitive before you build something more specialized.
runner = TaskRunner(
model="mlp_classifier",
task="classification",
epochs=PROFILE["epochs"],
batch_size=PROFILE["batch_size"],
label_key="louvain",
device="auto",
output_dir=str(OUTPUT_DIR),
)
runner.fit(adata)
metrics = runner.evaluate()
metrics
{'accuracy': 0.9343434343434344,
'macro_f1': 0.9204077302350909,
'weighted_f1': 0.9319851082871664,
'balanced_accuracy': 0.9036140509788966,
'macro_precision': 0.9468551399165805,
'macro_recall': 0.9036140509788966,
'weighted_precision': 0.9347937523014564,
'weighted_recall': 0.9343434343434344,
'cohen_kappa': 0.9107876267221211,
'mcc': 0.9115056195035095,
'confusion_matrix': [[51, 0, 0, 0, 0, 0, 0, 0],
[0, 72, 0, 0, 0, 0, 0, 0],
[1, 0, 166, 4, 0, 0, 0, 0],
[0, 0, 9, 35, 0, 0, 0, 4],
[0, 1, 0, 0, 5, 0, 0, 0],
[0, 4, 2, 0, 0, 17, 0, 0],
[0, 0, 0, 0, 0, 0, 2, 0],
[0, 0, 0, 1, 0, 0, 0, 22]],
'per_class_f1': [0.9902912621359223,
0.9664429530201343,
0.9540229885057471,
0.7954545454545454,
0.9090909090909091,
0.85,
1.0,
0.8979591836734694],
'per_class_precision': [0.9807692307692307,
0.935064935064935,
0.9378531073446328,
0.875,
1.0,
1.0,
1.0,
0.8461538461538461],
'per_class_recall': [1.0,
1.0,
0.9707602339181286,
0.7291666666666666,
0.8333333333333334,
0.7391304347826086,
1.0,
0.9565217391304348]}
Save a report and inspect the confusion matrix#
The notebook writes its report to artifacts/pbmc_classification/.
runner.save_report(OUTPUT_DIR / "report.md")
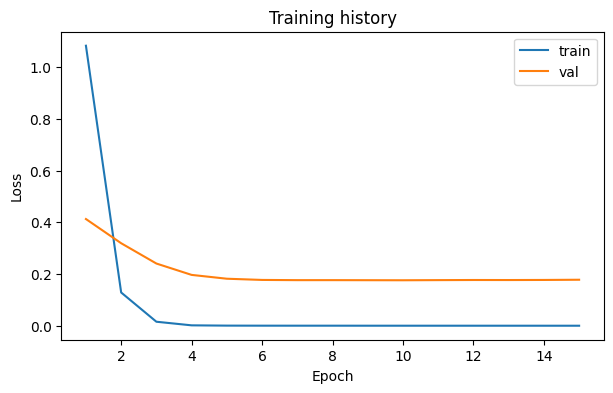
loss_fig, _ = runner.plot_losses()
loss_fig.savefig(OUTPUT_DIR / "loss_curve.png", dpi=150, bbox_inches="tight")
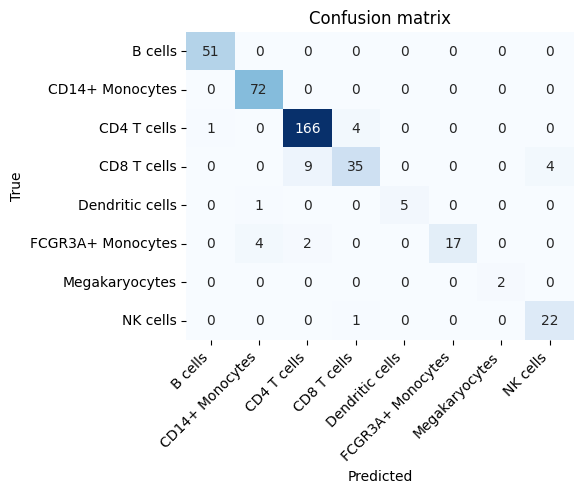
confusion_fig, _ = runner.plot_confusion_matrix()
confusion_fig.savefig(OUTPUT_DIR / "confusion_matrix.png", dpi=150, bbox_inches="tight")
display(confusion_fig)

Expected outputs#
After running the notebook, check:
accuracymacro_f1artifacts/pbmc_classification/report.mdartifacts/pbmc_classification/report.csvartifacts/pbmc_classification/loss_curve.pngartifacts/pbmc_classification/confusion_matrix.png
Interpretation checklist:
Treat this as a baseline classifier, not a production annotation system.
Read macro F1 before trusting accuracy if the class balance looks uneven.
Use the confusion matrix to see which PBMC groups the model mixes up most often.
If you want a stronger baseline before interpreting the results, switch the first config cell to TUTORIAL_PROFILE = "full" and rerun the notebook.
Recommended next tutorials:
PBMC model comparison for unsupervised baseline context
custom model extension if you want to build a task-specific classifier or encoder
output_paths = {
"report_md": str(OUTPUT_DIR / "report.md"),
"report_csv": str(OUTPUT_DIR / "report.csv"),
"loss_curve_png": str(OUTPUT_DIR / "loss_curve.png"),
"confusion_matrix_png": str(OUTPUT_DIR / "confusion_matrix.png"),
}
output_paths
{'report_md': 'artifacts/pbmc_classification/report.md',
'report_csv': 'artifacts/pbmc_classification/report.csv',
'loss_curve_png': 'artifacts/pbmc_classification/loss_curve.png',
'confusion_matrix_png': 'artifacts/pbmc_classification/confusion_matrix.png'}
